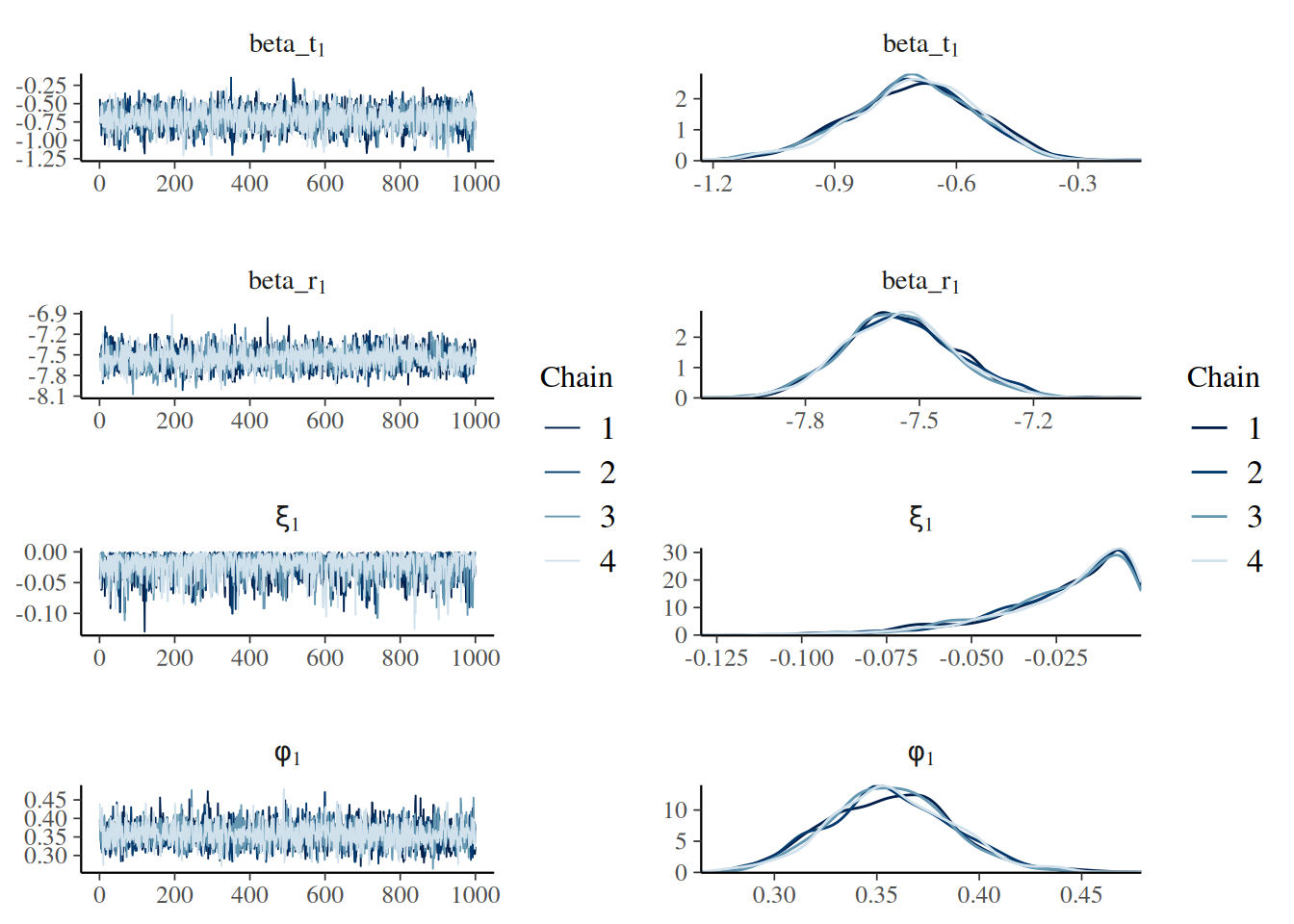
Provided the dataset is properly formatted, a simple model considering the existance of 8 age groups and a natural mortality rate of 0.25 can be fit as follows:
my_drm <-
fit_drm(.data = dat_train,
y_col = "dens",
time_col = "year",
site_col = "patch",
n_ages = 8,
m = 0.25,
seed = 2025)
Running MCMC with 4 sequential chains...
Chain 1 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 1 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 1 Exception: gamma_lpdf: Random variable is 0, but must be positive finite! (in '/tmp/Rtmp8LMTQe/model-2afb13976e39.stan', line 309, column 4 to column 54)
Chain 1 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 1 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 1 Iteration: 100 / 2000 [ 5%] (Warmup)
Chain 1 Iteration: 200 / 2000 [ 10%] (Warmup)
Chain 1 Iteration: 300 / 2000 [ 15%] (Warmup)
Chain 1 Iteration: 400 / 2000 [ 20%] (Warmup)
Chain 1 Iteration: 500 / 2000 [ 25%] (Warmup)
Chain 1 Iteration: 600 / 2000 [ 30%] (Warmup)
Chain 1 Iteration: 700 / 2000 [ 35%] (Warmup)
Chain 1 Iteration: 800 / 2000 [ 40%] (Warmup)
Chain 1 Iteration: 900 / 2000 [ 45%] (Warmup)
Chain 1 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 1 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 1 Iteration: 1100 / 2000 [ 55%] (Sampling)
Chain 1 Iteration: 1200 / 2000 [ 60%] (Sampling)
Chain 1 Iteration: 1300 / 2000 [ 65%] (Sampling)
Chain 1 Iteration: 1400 / 2000 [ 70%] (Sampling)
Chain 1 Iteration: 1500 / 2000 [ 75%] (Sampling)
Chain 1 Iteration: 1600 / 2000 [ 80%] (Sampling)
Chain 1 Iteration: 1700 / 2000 [ 85%] (Sampling)
Chain 1 Iteration: 1800 / 2000 [ 90%] (Sampling)
Chain 1 Iteration: 1900 / 2000 [ 95%] (Sampling)
Chain 1 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 1 finished in 2.5 seconds.
Chain 2 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 2 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 2 Exception: gamma_lpdf: Random variable is 0, but must be positive finite! (in '/tmp/Rtmp8LMTQe/model-2afb13976e39.stan', line 309, column 4 to column 54)
Chain 2 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 2 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 2 Iteration: 100 / 2000 [ 5%] (Warmup)
Chain 2 Iteration: 200 / 2000 [ 10%] (Warmup)
Chain 2 Iteration: 300 / 2000 [ 15%] (Warmup)
Chain 2 Iteration: 400 / 2000 [ 20%] (Warmup)
Chain 2 Iteration: 500 / 2000 [ 25%] (Warmup)
Chain 2 Iteration: 600 / 2000 [ 30%] (Warmup)
Chain 2 Iteration: 700 / 2000 [ 35%] (Warmup)
Chain 2 Iteration: 800 / 2000 [ 40%] (Warmup)
Chain 2 Iteration: 900 / 2000 [ 45%] (Warmup)
Chain 2 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 2 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 2 Iteration: 1100 / 2000 [ 55%] (Sampling)
Chain 2 Iteration: 1200 / 2000 [ 60%] (Sampling)
Chain 2 Iteration: 1300 / 2000 [ 65%] (Sampling)
Chain 2 Iteration: 1400 / 2000 [ 70%] (Sampling)
Chain 2 Iteration: 1500 / 2000 [ 75%] (Sampling)
Chain 2 Iteration: 1600 / 2000 [ 80%] (Sampling)
Chain 2 Iteration: 1700 / 2000 [ 85%] (Sampling)
Chain 2 Iteration: 1800 / 2000 [ 90%] (Sampling)
Chain 2 Iteration: 1900 / 2000 [ 95%] (Sampling)
Chain 2 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 2 finished in 2.5 seconds.
Chain 3 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: gamma_lpdf: Random variable is 0, but must be positive finite! (in '/tmp/Rtmp8LMTQe/model-2afb13976e39.stan', line 309, column 4 to column 54)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: gamma_lpdf: Random variable is 0, but must be positive finite! (in '/tmp/Rtmp8LMTQe/model-2afb13976e39.stan', line 309, column 4 to column 54)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: gamma_lpdf: Random variable is 0, but must be positive finite! (in '/tmp/Rtmp8LMTQe/model-2afb13976e39.stan', line 309, column 4 to column 54)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 3 Exception: gamma_lpdf: Random variable is 0, but must be positive finite! (in '/tmp/Rtmp8LMTQe/model-2afb13976e39.stan', line 309, column 4 to column 54)
Chain 3 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 3 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 3 Iteration: 100 / 2000 [ 5%] (Warmup)
Chain 3 Iteration: 200 / 2000 [ 10%] (Warmup)
Chain 3 Iteration: 300 / 2000 [ 15%] (Warmup)
Chain 3 Iteration: 400 / 2000 [ 20%] (Warmup)
Chain 3 Iteration: 500 / 2000 [ 25%] (Warmup)
Chain 3 Iteration: 600 / 2000 [ 30%] (Warmup)
Chain 3 Iteration: 700 / 2000 [ 35%] (Warmup)
Chain 3 Iteration: 800 / 2000 [ 40%] (Warmup)
Chain 3 Iteration: 900 / 2000 [ 45%] (Warmup)
Chain 3 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 3 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 3 Iteration: 1100 / 2000 [ 55%] (Sampling)
Chain 3 Iteration: 1200 / 2000 [ 60%] (Sampling)
Chain 3 Iteration: 1300 / 2000 [ 65%] (Sampling)
Chain 3 Iteration: 1400 / 2000 [ 70%] (Sampling)
Chain 3 Iteration: 1500 / 2000 [ 75%] (Sampling)
Chain 3 Iteration: 1600 / 2000 [ 80%] (Sampling)
Chain 3 Iteration: 1700 / 2000 [ 85%] (Sampling)
Chain 3 Iteration: 1800 / 2000 [ 90%] (Sampling)
Chain 3 Iteration: 1900 / 2000 [ 95%] (Sampling)
Chain 3 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 3 finished in 2.6 seconds.
Chain 4 Iteration: 1 / 2000 [ 0%] (Warmup)
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: gamma_lpdf: Random variable is 0, but must be positive finite! (in '/tmp/Rtmp8LMTQe/model-2afb13976e39.stan', line 309, column 4 to column 54)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: gamma_lpdf: Random variable is 0, but must be positive finite! (in '/tmp/Rtmp8LMTQe/model-2afb13976e39.stan', line 309, column 4 to column 54)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: gamma_lpdf: Random variable is 0, but must be positive finite! (in '/tmp/Rtmp8LMTQe/model-2afb13976e39.stan', line 309, column 4 to column 54)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4 Informational Message: The current Metropolis proposal is about to be rejected because of the following issue:
Chain 4 Exception: gamma_lpdf: Random variable is 0, but must be positive finite! (in '/tmp/Rtmp8LMTQe/model-2afb13976e39.stan', line 309, column 4 to column 54)
Chain 4 If this warning occurs sporadically, such as for highly constrained variable types like covariance matrices, then the sampler is fine,
Chain 4 but if this warning occurs often then your model may be either severely ill-conditioned or misspecified.
Chain 4 Iteration: 100 / 2000 [ 5%] (Warmup)
Chain 4 Iteration: 200 / 2000 [ 10%] (Warmup)
Chain 4 Iteration: 300 / 2000 [ 15%] (Warmup)
Chain 4 Iteration: 400 / 2000 [ 20%] (Warmup)
Chain 4 Iteration: 500 / 2000 [ 25%] (Warmup)
Chain 4 Iteration: 600 / 2000 [ 30%] (Warmup)
Chain 4 Iteration: 700 / 2000 [ 35%] (Warmup)
Chain 4 Iteration: 800 / 2000 [ 40%] (Warmup)
Chain 4 Iteration: 900 / 2000 [ 45%] (Warmup)
Chain 4 Iteration: 1000 / 2000 [ 50%] (Warmup)
Chain 4 Iteration: 1001 / 2000 [ 50%] (Sampling)
Chain 4 Iteration: 1100 / 2000 [ 55%] (Sampling)
Chain 4 Iteration: 1200 / 2000 [ 60%] (Sampling)
Chain 4 Iteration: 1300 / 2000 [ 65%] (Sampling)
Chain 4 Iteration: 1400 / 2000 [ 70%] (Sampling)
Chain 4 Iteration: 1500 / 2000 [ 75%] (Sampling)
Chain 4 Iteration: 1600 / 2000 [ 80%] (Sampling)
Chain 4 Iteration: 1700 / 2000 [ 85%] (Sampling)
Chain 4 Iteration: 1800 / 2000 [ 90%] (Sampling)
Chain 4 Iteration: 1900 / 2000 [ 95%] (Sampling)
Chain 4 Iteration: 2000 / 2000 [100%] (Sampling)
Chain 4 finished in 2.8 seconds.
All 4 chains finished successfully.
Mean chain execution time: 2.6 seconds.
Total execution time: 10.8 seconds.
Warning: 7 of 4000 (0.0%) transitions ended with a divergence.
See https://mc-stan.org/misc/warnings for details.